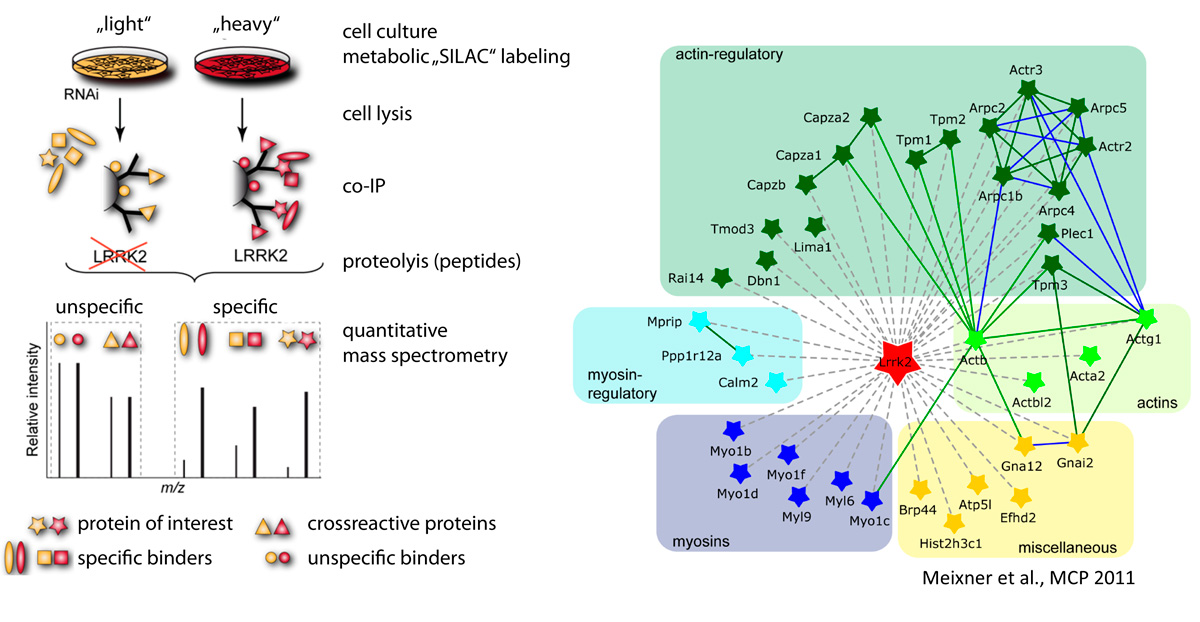
The workgroup focuses on the analysis of proteins and functional protein networks associated with pathogenic gene variants causing Parkinson’s disease (PD). By systematically following the ‘guilt-by-association principle’ we aim at defining functional protein networks altered by disease to identify mutation-specific molecular pathomechanisms. Towards this goal, we are combining affinity-based purification with quantitative mass spectrometry (AP-MS) to identify protein-protein networks associated with proteins mutated in familial forms of PD. In addition, complementary approaches, such as BioID-based proximity labelling, are used to map functional protein complexes within intact cells.
In addition, mass spectrometric mapping is used to identify disease-specific posttranslational protein modification patterns (i.e. protein phosphorylation, acetylation and ubiquitination)
Taking advantage of the multidisciplinary environment at the DZNE, the findings are subsequently validated by functional analyses in cell systems such as patient-derived induced pluripotent stem cells (iPS).
Given that proteins mutated in neurodegenerative diseases, such as LRRK2, and proteins of associated functional networks might be suitable drug-targets for disease treatment we are additionally studying structure-function relationships by structural proteomics.
By learning from mutation-specific molecular signatures as well as perturbations of functional protein networks and posttranslational modification patterns we aim to define rational biomarkers relevant for diagnostics and drug development.
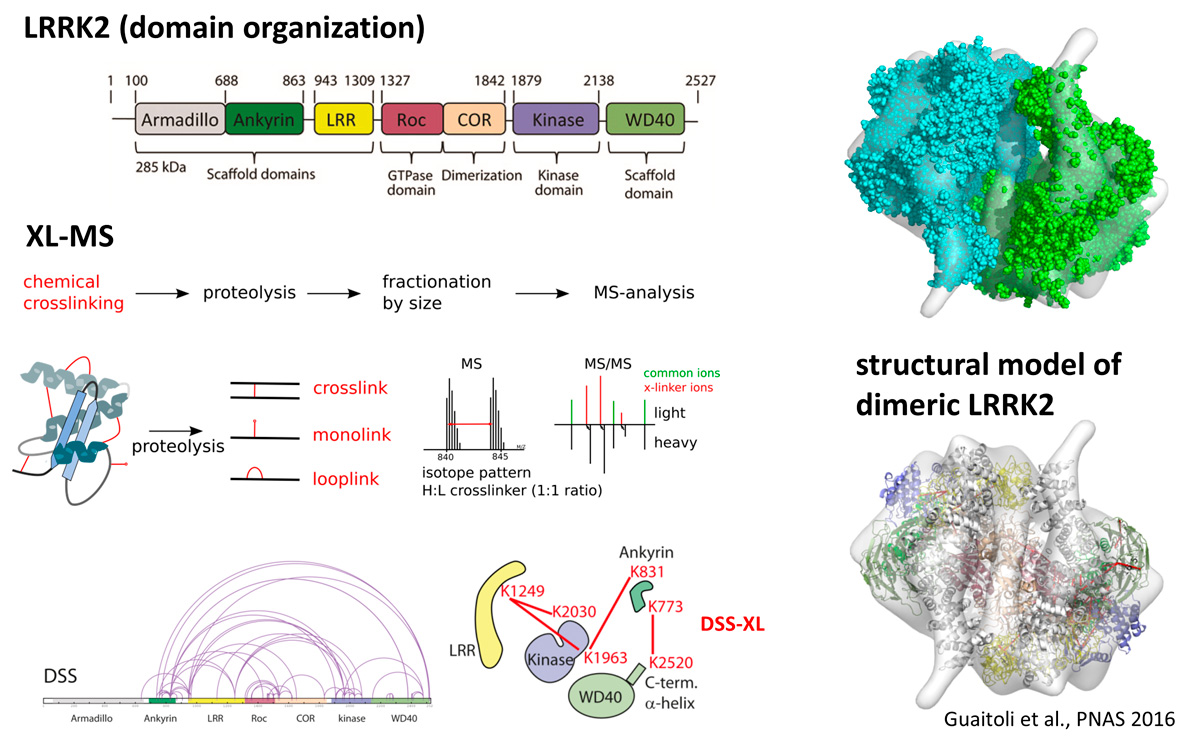
Given that mutations in the LRRK2 gene are the most common reason of familial forms of PD with known genetic cause, a strong focus is laid on the systematic analysis of the gene product, the Leucine-rich repeat kinase 2 protein. Since PD-associated LRRK2 variants show decreased GTPase and increased kinase activity, LRRK2 is also a promising drug-target. To allow a mechanistic understanding at a molecular and atomic level and to provide a foundation for rational drug design, the availability of a protein structure is very important. A large and flexible protein, such as LRRK2 it is however very hard to crystalize.
By integrating multiple experimental inputs provided by chemical cross-linking, small-angle X-ray scattering, and a negative-stain
EM map, we nevertheless could recently build a first structural model of the full-length LRRK2 dimer (Guaitoli et al., 2016). The hybrid model reveals a compact folding of the LRRK2 dimer with multiple domain–domain interactions that might be involved in the regulation of LRRK2 enzymatic properties.
Our group now focusses on the refinement of the model also addressing the in-depth biochemical analysis of the LRRK2 G-domain and its regulation by adjacent protein domains.
Funding:
- Young investigator group funding as part of iMED program of the Helmholtz Association
- From structure and function to allosteric targeting of LRRK2-mediated PD; Michael J Fox Foundation LRRK2 Biology (2018)